Perry R. Bass Marine Fisheries Research Station
Research is a critical asset for managing the Texas fishery. Among other applications, it is very beneficial for improving the current stocking methodology and developing procedures for new goals, such as successful mass stocking of southern flounder. To these ends, the staff at Perry R. Bass Research Station is divided into two groups: life history and genetics. The life history biologists examine age, growth, and reproductive characteristics of marine organisms. The geneticists study genetic variances between different populations of the same species. Together, the research from these two specialties provides information for the responsible management of Texas's marine resources.(1)
Life History
The goal of the life history program is to collect information on fish populations that are not adequately explored using routine monitoring data. The main areas of life history research being conducted at Perry R. Bass are reproduction, age determination, and temperature tolerance. Age determination is significant in "estimating growth and mortality rates, population age structure, and other parameters needed for understanding the population dynamics of important Gulf of Mexico fish stocks and their response to natural phenomena and exploitation."(2) This information can then be translated into regulations that protect the fishery. For instance, spotted seatrout females are sexually mature at 13" (approx. 1.5 years). Therefore, the minimum size limit is 15"; this way, the trout has a chance to spawn (and contribute to the continuation of her species) before she is potentially taken out of the reproductive pool, permanently. Redfish are another story. With this species extended juvenile period, they don't reach sexual maturity for 4 or 5 years, and once they do, they leave the bays! If the same strategy to regulate trout was used to regulate redfish, the bay fishery for them would essentially cease to exist. In response, and with the life history information for redfish, a slot limit and three fish bag was imposed: only redfish 20-28" are fair game (and one oversized red per day). This allows a strong up-and-coming base and also protects a majority of sexually mature adults.(3) The whole point of this, of course, is to not take at a faster rate than the population can replenish. We don't want to end up with just one truffula seed.
The main vehicle for age determination is the otolith (fish ear bone). Though not all fish can be accurately aged by their otoliths, most along the Texas coast can, and it's important to adopt "standardized ageing criteria for each species [in order to] provide comparable information necessary for age structured stock assessments at state and regional levels."(2) Kind of like how the SATs provide colleges with a level playing field (in theory) for evaluating applicants. Reading otoliths is like counting tree rings. For most fish, one opaque and one translucent ring form on the otolith per year, so the number of opaque rings is equal to the fish's age in years. For fish that can't be aged via their otoliths, alternative means include spines, finrays, vertebrae, and scales. Scientists can read more than just age on an otolith, though. They can often find "a cumulative historical record of changes in climate, nutrition, hydrographic environment, and other ecological parameters."(2) It's like a CD-ROM of the fish's life. For some long-lived drum species, the otolith can be tested for radiocarbon resulting from atomic bomb testing in the 1950s and 1960s. There's even a way for scientists to evaluate the environmental conditions of long extinct populations.(2) Yes, we know how fish partied in the Pleistocene.
The original reasoning behind testing temperature tolerances was to discover at what age young hatchery fish could be safely stocked into growout ponds. Typically, the staff at Perry R. Bass studies reds, seatrout, and flounder (those being the principle stock enhancement fish) from two weeks to forty-five days old (depending on species and study goals). Using an environmental chamber, they can mimic natural temperature changes of any time of the year. A study was recently completed on young, post-metamorphose flounder (the age often stocked into growout ponds) to test their resilience to freezing temperatures. Most of the flounder in this study (three tests, 30 fish each test) made it through an extended (south Texas) freeze of 48 hours just fine, proving that stocking this age group during the winter is not a problem. That's great for a (semi)controlled environment, but results from these tests can also pertain to wild stock. For example, fish become more sluggish during cold temperatures, and some fish will move into deeper water, such as the Intracoastal Waterway, when the temperature drops. As they are slower to react to passing barges, and there are more of them in the ICW, mortality rates from barge run-ins increase during those periods of cold temperature. However, if we know the minimum temperature at which the fish can safely avoid the barges (provided they recognize the barges as a danger in the first place), the barge operators can be made aware of when their runs through the ICW are most harmful, and some may be able to reschedule or delay their runs to accommodate.(3)
Genetics
The function of the population genetics lab at Perry R. Bass, according to the TPWD website, is to "apply genetic techniques to answer questions raised by fishery managers of the Coastal Fishery Division of Texas Parks and Wildlife Department."(1) This is where the determination of genetic variances comes in. It is important to determine if multiple groups of the same species interbreed or not. If they don't, they should be managed separately. If they do, they should be managed as one group. That is the chief value of genetics in management decisions. The stock enhancement program also benefits from this branch of research. It ensures that hatchery-cultured fish are released into the same genetic stock from whence their parents were drafted. For example, flounder from different bay systems are genetically distinct, so the offspring of flounder from Matagorda Bay would only be stocked in Matagorda Bay. Seatrout are generally classified in one of three genetic populations: those from the upper coast bays, middle coast bay system, and lower coast system.(3) Since redfish spawn offshore (as opposed to seatrout and flounder who spawn inshore), there is little to no genetic variance of the wild stock along the Texas coast (but those ones from Florida just can't keep their Disney hidden).
This research is primarily conducted to preserve the natural state of wild stocks. Maybe nothing would happen if flounder from Matagorda Bay were released in the Lower Laguna, but there's no reason to take that chance when there are so many historical examples of environmental disruption after human interaction (carp in the Great Lakes, smallmouth bass in the Guadalupe bass habitat, etc). For this reason, there are also theoretical "what if" studies conducted. Cobia are offshore spawners, but what if someone cultured cobia in a bay system? How would that affect offshore population? Oysters also have genetically distinct populations within the bays. What if someone wanted to start an oyster farm or transplant a population? How would that affect the existing populations? These situations might never happen, but it's good to know what the effects would be if they did.(3)
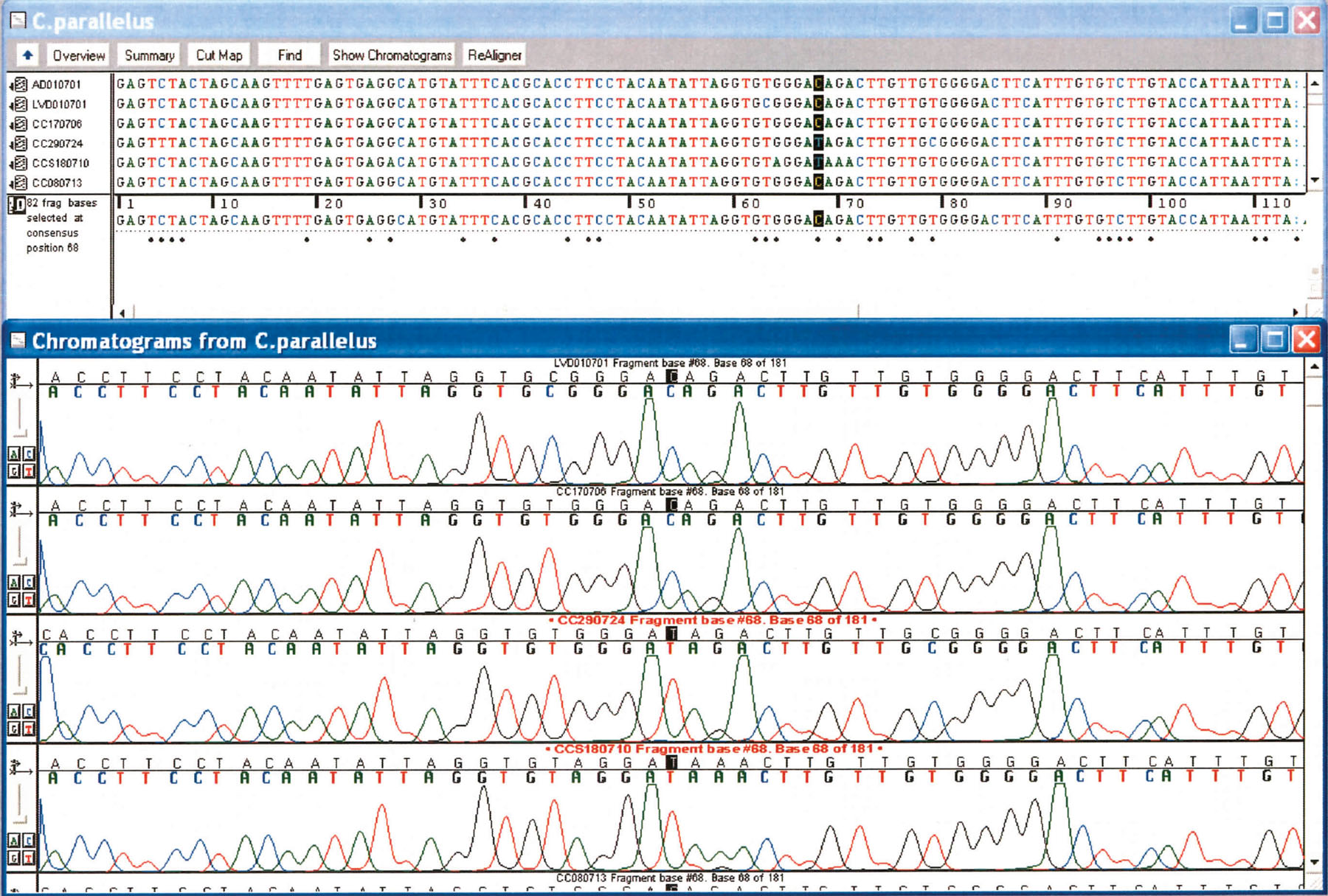
The way genetic variances are studied is through DNA sequencing. From fin clips (and using sophisticated government equipment), the biologist isolates a particular DNA sequence from a single fish and then compares that to sequences obtained from other fish of the same species. The optimal sequence is the one that has the most potential differences between individual fishes. Through past research, the choice is fairly predictable, though there is still a small amount of trial and error involved sometimes. Depending on the study, 30-40 individual finclips (from separate fish) are needed per system (for however many systems are being compared) to have enough data for accurate conclusions. Typically, there is a 1-2% difference between sequences. If there is a 5% difference (or higher), there might be potential for considering the possibility of subspecies.(3)
Questions regarding fishery management are always available for genetic problem solving. There is an ongoing study into Gulf menhaden to determine whether or not there are distinct genetic populations along the coast and whether or not the commercial fishery might be stressing one particular population. Early results say no to the former (and therefore, no to the latter), but there haven't been enough fish sampled to be sure.(3)
Fish seem to sustain a never-ending field of inquiry. For every puzzle you think you've completed, there always seems to be an extra piece leading you to some other anomaly.
There is nothing like looking, if you want to find something. You certainly usually find something, if you look, but it is not always quite the something you were after. ~J.R.R. Tolkien
Works Cited
(1) "Perry R. Bass Marine Fisheries Research Center," Texas Parks & Wildlife Department, 10 January 2013 <http://www.tpwd.state.tx.us/fishboat/fish/management/hatcheries/prb.phtml>.
(2) "A Practical Handbook for Determining the Ages of Gulf of Mexico Fishes: Second Edition," (Gulf States Marine Fisheries Division, pp. 10-15, 2009)
(3) The excellent staff at Perry R. Bass Marine Fisheries Research Station


